

METAflow is a novel “augmented intelligence”-driven analysis solution for flow- and mass- cytometry
Our proprietary AI reduces the need for technical expertise to interpret data,
provides a flexible environment for reviewing your results,
and includes an intuitive dashboard revealing outliers and linking the entirety of your analysis workflow
To see how you can achieve objective and reproducible results in record time,
click the link below for more information.
Want to receive updates and news about our latest versions of METAflow:
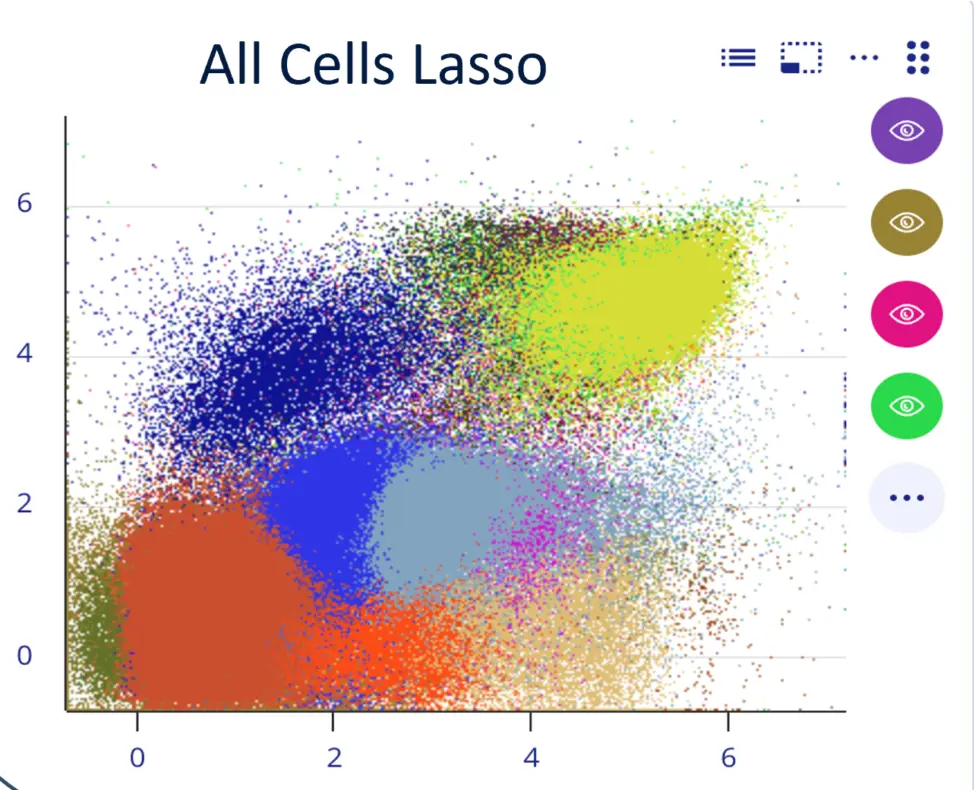
Facing analysis bottlenecks with too many cells, files, parameters, and limited expertise?
METAflow is a cloud-based computing platform designed to do the heavy lifting and present the results in an easily interpretable interface. You define the biology, let METAflow handle the analysis.
No more bottlenecks, just streamlined efficiency.
Struggling with reproducibility, where lack of technical expertise results in significant variability and challenges the interpretation of crucial quantitative results?
METAflow reduces technical barriers by providing an accurate representation of the data, without the need for second-guessing gate boundaries, double-counting of cells, or cross-population contamination.
Achieve accurate results from the start, consistently.
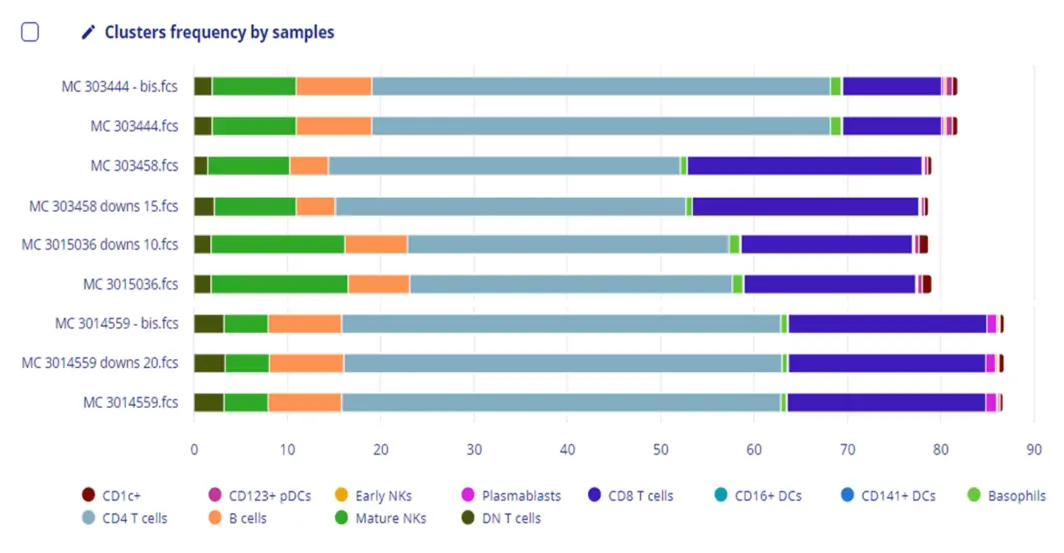
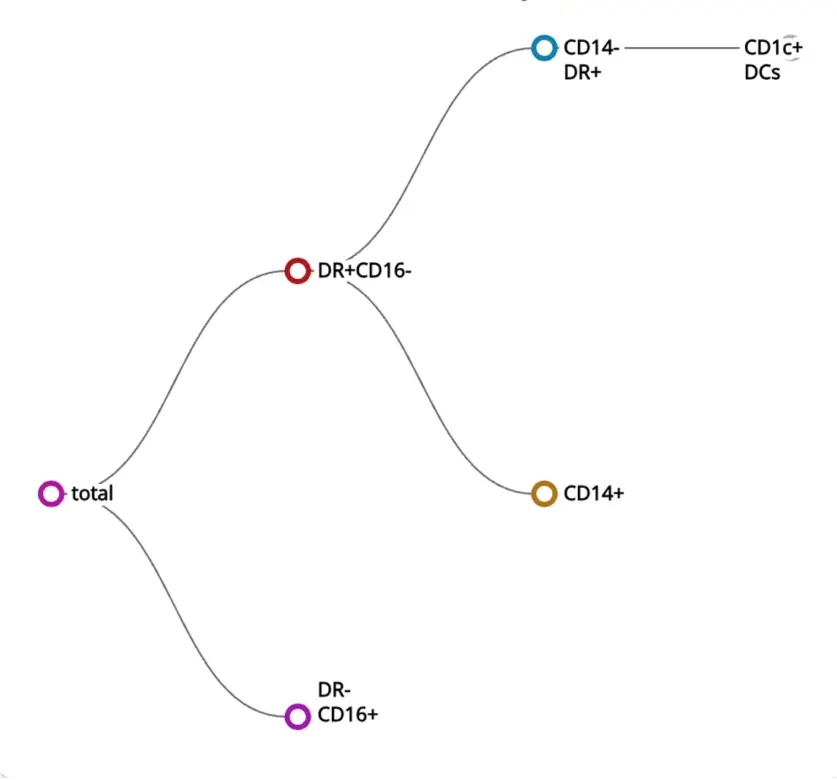
Encountering challenges with the heavy burden of understanding the underlying math or computer science behind automated solutions?
The built-in AI handles the technical components for you. The integrated AI automatically adjusts the technical components of each algorithm, so that you can focus on the biology.
METAflow is your data analysis co-pilot.

Alan Saluk
Senior Scientific Director
at Scripps Research

Raïf Yucel
Head of Centre of Cytomics
at University of Exeter

“While hardware and reagents have allowed for high-dimensional data sets in flow cytometry, the majority of our analytical techniques are shockingly antiquated in two dimensions. METAflow is a gateway for modern high-dimensional data analyses”

“While hardware and reagents have allowed for high-dimensional data sets in flow cytometry, the majority of our analytical techniques are shockingly antiquated in two dimensions. METAflow is a gateway for modern high-dimensional data analyses”

“AI-based software tools are needed to obtain more unbiased scientific results. Uniquely, METAflow opens an innovative roadmap to analyze a high-dimensional dataset with infinite potential. Especially, METAflow lays a promising basis to develop digital clinical cytometry research and diagnostic approaches”

This project has received funding under European Union’s Horizon Europe programme, under Grant Agreement n. 190166506
Pharma
High throughput screening, data security
Biotech
Assay development and patient monitoring
Academic
Discovery tool for complex data
CROs
One tool to unite them all
Clinical Labs
21 CFR P11 compliance
High throughput screening, data security
Assay development and patient monitoring
Discovery tool for complex data
One tool to unite them all
21 CFR P11 compliance
